Plant-associated microbial communities can profoundly affect plant health and success, and research is still uncovering factors driving the assembly of these communities. Here, we examine how geography versus host species affects microbial community structure and differential abundances of individual taxa. We use metabarcoding to characterize the bacteria and eukaryotes associated with five, often co-occurring species of Sarracenia pitcher plants (Sarraceniaceae) and three natural hybrids along the longitudinal gradient of the U.S. Gulf Coast, as well as samples from S. purpurea in Massachusetts. To tease apart the effects of geography versus host species, we focus first on sites with co-occurring species and then on species located across different sites. Our analyses show that bacterial and eukaryotic community structures are clearly and consistently influenced by host species identity, with geographic factors also playing a role. Naturally occurring hybrids appear to also host unique communities, which are in some ways intermediate between their parent species. We see significant effects of geography (site and longitude), but these generally explain less of the variation among pitcher communities. Overall, in Sarracenia pitchers, host plant phenotype significantly affects the pitcher microbiomes and other associated organisms.
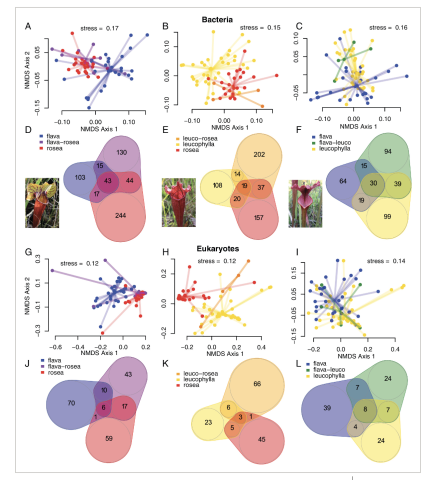
Beta diversity and Venn diagrams of bacterial and eukaryotic communities associated with hybrids (see inset photos for examples). NMDS plots using weighted UniFrac distances of bacterial (A–C) and eukaryotic (G–I) communities associated with natural hybrids and their parent species: S. rosea, S. leucophylla and S. flava. Venn diagrams show bacterial (D–F) and eukaryotic (J–L) ASVs normalized by sample size that are shared between natural hybrids and their parent species.
| GEM3 author(s) | |
| Year published |
2022
|
| Journal |
Environmental Microbiology
|
| DOI/URL | |
| Keywords |
Bioinformatics
Ecology
Genomics
Geography
Population Genetics
|
| GEM3 component |
Mechanisms
|
| Mentions grant |
Yes
|